《好想睡覺的小兔子》(The Rabbit Who Wants to Fall Asleep)這本書的中文版最近要在台灣上市了,很多有小小孩的家長對於這件事情非常關心,很想瞭解是否真的能達到效果?對於嬰幼兒的睡眠能否找到一勞永逸的方法?

其實,這本書的英文版本去年就已經出版,當時曾在國外造成一股風潮。但是很多有實驗精神的心理學家,也試著在自己的小孩身上試用,但效果似乎不是很好。(請見:想睡覺的兔子讓你家寶寶入睡了嗎?)
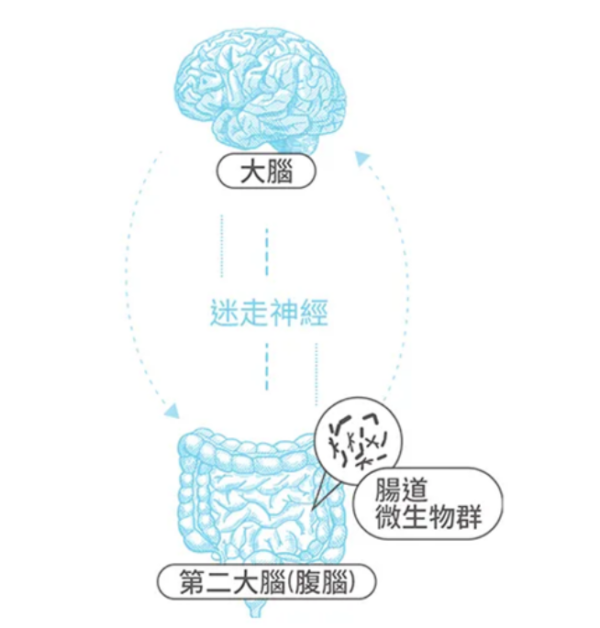
個人瞭解了英文版本(中文版本還未到手),發現它是使用放鬆的概念,並企圖營造出催眠的感覺(放鬆一事是催眠的基礎,不能放鬆就不可能被催眠)。裡面所使用的語氣與詞彙,都是想造成睡眠的氛圍。但這樣的概念,其實出自於睡眠儀式。大人其實只要想辦法營造睡眠氣氛,一樣可達到相同效果。

像我家的雙胞胎用這樣的方式肯定無效,因為我們睡前不講故事的。由神經科學可知,講故事這件事牽涉到認知學習。如果我們想講故事,都是盡量在清醒的狀態,這樣所說的故事才有可能觸動大腦學習。我們的作法只是讓睡前的整個流程盡量簡化,並且盡可能在15分鐘內完成。我們是這樣做的:先泡奶、喝奶,之後上廁所、包尿布、清口腔,最後全家人一起上床睡覺。他們當然不會一下子就睡著,但是滾來滾去大約十五分鐘內就會進入夢鄉。這與大部分的睡眠科學研究相符合,大部分的人都是十五到三十分鐘內睡著(見延伸閱讀)。
對於睡眠儀式有疑慮的人,會覺得這樣流程不容易建立,甚至弄得不好整個過程會長達一兩小時。但這真的是誤解,如果放學回家就要開始進行睡眠儀式,這完全是不懂行為理論所導致。我們先停下來思考一下大人怎麼睡覺的,就會發現有疑問的人完全搞錯方向了。成年人想睡的時間,與生理時鐘配合。有的人早一些大約十點就想睡了,有的人晚一些,大約要十二點才想睡覺。每個人覺得睡得夠飽的時間也不同,落差在四到十小時之間(若以歷史上知名的人物來看,拿破崙只需要睡四小時,但是愛因斯坦卻要睡到十小時以上)。因此,大人在想睡覺的時候,睡眠儀式很簡短,準備一下就睡了,不需要拖到一、兩個小時(註一)。

所以,回過頭來看嬰幼兒的睡眠也是如此。小孩想睡的時間到了,生理時鐘就是這樣,不可能撐太久,幾乎都會很快就睡著。因此睡眠儀式只要睡覺前幾個步驟就行了,而不是遙遙無盡期。若有一兩天失敗,這是很正常的,因為孩子們不是機器人,關了機就可以直接放倒在地上。每個孩子會因為當天的活動量、誤吃咖啡因相關食物、使用藥物、有無照到太陽(註二),而略有不同的。我家的雙胞胎也曾經喝到茶,而搞到十一、二點才睡著。
總之,千萬不要誤信單一方法或書籍可以永久地解決所有問題。這不只不科學,也容易造成巨大的誤會,以為書買回家照做就會成功。不管是大人或小孩,都是有機體,我們要找尋的是合適的方式,而非人云亦云。
- 註一:如果每天真的要拖一兩個小時才能睡著,每星期有三天有這樣的情形,持續了三個月。那很有可能是睡眠節律有問題,應該要尋求醫療專業協助。
- 註二:太陽光可以幫忙校正松果體,讓松果體正常釋出褪黑激素。褪黑激素可以幫助我們入眠。
延伸閱讀:
參考文獻:
- American Psychiatric Association: Diagnostic and Statistical Manual of Mental Disorders,
5th edition. Arlington, VA., American Psychiatric Association, 2013. - Mindell JA, Kuhn B, Lewin DS et al. Behavioral treatment of bedtime problems and night
wakings in infants and young children. SLEEP 2006;29(10):1263-1276.
本文轉載自作者部落格暗夜浮動月黃昏